Large-scale human data show that a single antibiotic course can leave a measurable microbial footprint years later, raising important questions about long-term health effects and prescribing practices.
Study: Individual-level prescription data from 14,979 individuals link antibiotic use and gut microbiome composition.. Image Credit: Marevgena/Shutterstock
In a recent study published in the journal naturopathyResearchers found that the effects of antibiotics on the gut microbiome may last much longer than expected. By combining Swedish prescription registry data with fecal metagenomic profiles of nearly 15,000 adults in several population-based cohorts, the study showed that a single antibiotic course taken four to eight years earlier was associated with lasting differences in microbiome composition.
These findings highlight the potential for long-term microbiome disruption and underscore the need for careful antibiotic use, as even routine prescriptions can have lasting health effects.
Gut Microbiome Disruption and Long-Term Disease Risk
Disruption of the gut microbial ecosystem is being recognized as a major factor in a wide range of health conditions, including cardiometabolic disease, autoimmune disorders, and colorectal cancer. Antibiotics are a major driver of these imbalances, and frequent or long-term use is linked to higher risks of obesity, type 2 diabetes, and heart disease.
Small intervention studies have shown rapid declines in microbial diversity and shifts in dominant bacterial groups after treatment. However, these findings largely reflect short-term effects. As antibiotic use remains widespread, understanding long-term effects on gut microbial communities has become an important research priority, although large-scale robust evidence is limited.
Population study design and microbiome analysis methods
In this population-level study of 14,979 individuals, researchers examined the effect of oral antibiotic use over the past eight years on gut microbial composition. They combined prescription data from the Swedish National Prescribed Drug Register (NPDR) with fecal metagenomic data from three groups: the Cardiopulmonary Bioimage Study (SCAPIS), the Swedish Infrastructure for Medical Population-Based Life-Course and Environmental Research (SIMPLAR), and the Malmö Offspring Study (MOS).
The team assessed antibiotic exposure over three time periods: within 1 year, 1 to 4 years, and 4 to 8 years before sampling. Participants with recent antibiotic use, inflammatory bowel disease, or chronic pulmonary disease were excluded. Fecal samples underwent deep shotgun metagenomic sequencing, and microbial diversity was quantified using indices such as the Shannon index and the inverse Simpson index. Taxonomic classification was performed using the Genome Taxonomy Database.
Multivariate regression models assessed the relationship between antibiotic exposure and microbiome diversity while adjusting for confounders including age, sex, body mass index (BMI), smoking status, education, and co-morbidities. The model also accounts for medications such as proton-pump inhibitors (PPIs), statins, metformin, beta-blockers, antipsychotics, and selective serotonin reuptake inhibitors (SSRIs).
Sensitivity analyzes tested alternative exclusion criteria, while spline-based models explored microbiome recovery over time. Subgroup analyzes were conducted based on age and sex.
Persistent microbiome changes after antibiotic exposure
Antibiotic use was common, with approximately 70–74% of participants having at least one prescription in the past eight years. The broad-spectrum penicillins, penicillin V, and tetracycline were most commonly used.
The greatest reduction in microbial diversity occurred within 1 year of antibiotic use. However, significant associations persisted for exposures one to four years earlier and even four to eight years earlier. Each additional course was associated with a further decline in diversity. Notably, a single course taken years earlier was associated with long-term differences in the microbiome.
Clindamycin, fluoroquinolones, and flucloxacillin showed the strongest and most persistent effects. These antibiotics were associated with changes in the relative abundance of microbial species by approximately 10–15%. Beneficial species like alistipes communis And Odoribacter splanchnicus rejected. In contrast, species associated with metabolic risk, including Ruminococcus gnavus, Flavoniferous PlautyAnd agerthella lentagrew in abundance. Penicillin V showed relatively little effect.
Microbiome recovery was fastest within the first 2 years after antibiotic use but slowed thereafter and was often incomplete. Recovery varies by antibiotic class, gender, and age, suggesting that microbiome resilience varies among individuals. Sensitivity analysis confirmed the robustness of these findings.
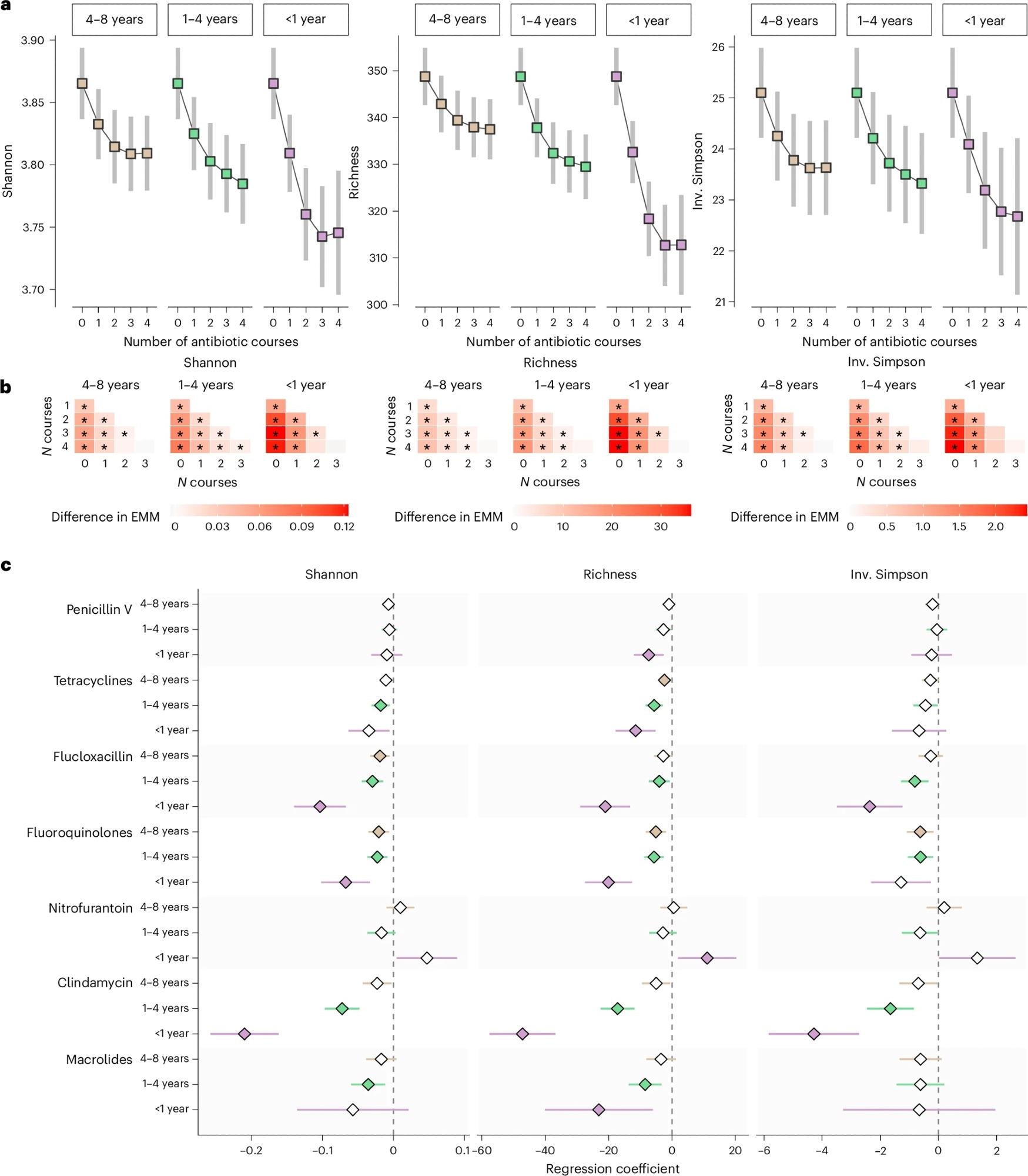
AMicrobiome diversity metrics (Shannon index, species richness and inverse (inv.) Simpson index) for each additional course of any antibiotic at 4–8 years, 1–4 years and <1 year before stool sampling. Estimated marginal means of heterogeneity (EMM) were obtained using regression models that included restricted cubic splines and adjustment for age, sex, smoking, education, country of birth, site-specific analysis plate, BMI, Charlson comorbidity index, polypharmacy, and use of PPIs, metformin, SSRIs, statins, beta-blockers, and antipsychotics (n = 14,974). Was modeled. Squares represent EMM, and 95% stop confidence intervals. bPairwise differences in EMM of microbiome diversity based on number of previous antibiotic courses. Stars indicate significant differences (FDR < 5%). CThe relationship between antibiotic use in the 8 years before fecal sampling and gut microbiome species diversity was examined using regression models adjusted for the same covariates as above in the three groups (SCAPIS, SIMPLAR, MOS, total n = 14,974), followed by meta-analyses of regression coefficients. The x axis and diamonds display meta-analysis regression coefficients; Error bars represent 95% confidence intervals. Filled symbols indicate statistically significant associations (FDR < 5%). At least one antibiotic has been shown to have such a relationship.
Implications for antibiotic stewardship and future research
The findings suggest that some antibiotics may have long-lasting effects on the gut microbiome, persisting for more than four years. These results support more prudent antibiotic prescribing and management strategies that minimize microbiome disruption.
However, the study remains observational and cannot establish causation. More research is needed to understand long-term health consequences, including associations with chronic disease. The findings cannot be generalized beyond outpatient settings in Sweden and total antibiotic exposure may be underestimated due to missing hospital or international prescription data.
Journal Reference:
- Baldanzi, G. et al. (2026). Individual-level prescription data from 14,979 individuals link antibiotic use and gut microbiome composition. Naturopathy, 1-11. DOI: 10.1038/s41591-026-04284-y, https://www.nature.com/articles/s41591-026-04284-y

